Bayesian meta-analysis assuming that the effect size \(d\) varies across studies with standard deviation \(\tau\) (i.e., a random-effects model).
meta_random(
y,
SE,
labels,
data,
d = prior("cauchy", c(location = 0, scale = 0.707)),
tau = prior("invgamma", c(shape = 1, scale = 0.15)),
rscale_contin = 0.5,
rscale_discrete = 0.707,
centering = TRUE,
logml = "integrate",
summarize = "stan",
ci = 0.95,
rel.tol = .Machine$double.eps^0.3,
logml_iter = 5000,
silent_stan = TRUE,
...
)Arguments
- y
effect size per study. Can be provided as (1) a numeric vector, (2) the quoted or unquoted name of the variable in
data, or (3) aformulato include discrete or continuous moderator variables.- SE
standard error of effect size for each study. Can be a numeric vector or the quoted or unquoted name of the variable in
data- labels
optional: character values with study labels. Can be a character vector or the quoted or unquoted name of the variable in
data- data
data frame containing the variables for effect size
y, standard errorSE,labels, and moderators per study.- d
priordistribution on the average effect sized. The prior probability density function is defined viaprior.- tau
priordistribution on the between-study heterogeneitytau(i.e., the standard deviation of the study effect sizesdstudyin a random-effects meta-analysis. A (nonnegative) prior probability density function is defined viaprior.- rscale_contin
scale parameter of the JZS prior for the continuous covariates.
- rscale_discrete
scale parameter of the JZS prior for discrete moderators.
- centering
whether continuous moderators are centered.
- logml
how to estimate the log-marginal likelihood: either by numerical integration (
"integrate") or by bridge sampling using MCMC/Stan samples ("stan"). To obtain high precision withlogml="stan", many MCMC samples are required (e.g.,logml_iter=10000, warmup=1000).- summarize
how to estimate parameter summaries (mean, median, SD, etc.): Either by numerical integration (
summarize = "integrate") or based on MCMC/Stan samples (summarize = "stan").- ci
probability for the credibility/highest-density intervals.
- rel.tol
relative tolerance used for numerical integration using
integrate. Userel.tol=.Machine$double.epsfor maximal precision (however, this might be slow).- logml_iter
number of iterations (per chain) from the posterior distribution of
dandtau. The samples are used for computing the marginal likelihood of the random-effects model with bridge sampling (iflogml="stan") and for obtaining parameter estimates (ifsummarize="stan"). Note that the argumentiter=2000controls the number of iterations for estimation of the random-effect parameters per study in random-effects meta-analysis.- silent_stan
whether to suppress the Stan progress bar.
- ...
further arguments passed to
rstan::sampling(seestanmodel-method-sampling). Relevant MCMC settings concern the number of warmup samples that are discarded (warmup=500), the total number of iterations per chain (iter=2000), the number of MCMC chains (chains=4), whether multiple cores should be used (cores=4), and control arguments that make the sampling in Stan more robust, for instance:control=list(adapt_delta=.97).
Examples
# \donttest{
### Bayesian Random-Effects Meta-Analysis (H1: d>0)
data(towels)
set.seed(123)
mr <- meta_random(logOR, SE, study,
data = towels,
d = prior("norm", c(mean = 0, sd = .3), lower = 0),
tau = prior("invgamma", c(shape = 1, scale = 0.15))
)
mr
#> ### Bayesian Random-Effects Meta-Analysis ###
#> Prior on d: 'norm' (mean=0, sd=0.3) truncated to the interval [0,Inf].
#> Prior on tau: 'invgamma' (shape=1, scale=0.15) with support on the interval [0,Inf].
#>
#> # Bayes factors:
#> (denominator)
#> (numerator) random_H0 random_H1
#> random_H0 1.00 0.262
#> random_H1 3.81 1.000
#>
#> # Posterior summary statistics of random-effects model:
#> mean sd 2.5% 50% 97.5% hpd95_lower hpd95_upper n_eff Rhat
#> d 0.196 0.088 0.032 0.194 0.373 0.015 0.349 5694.0 1
#> tau 0.129 0.087 0.033 0.106 0.355 0.022 0.299 4694.1 1
plot_posterior(mr)
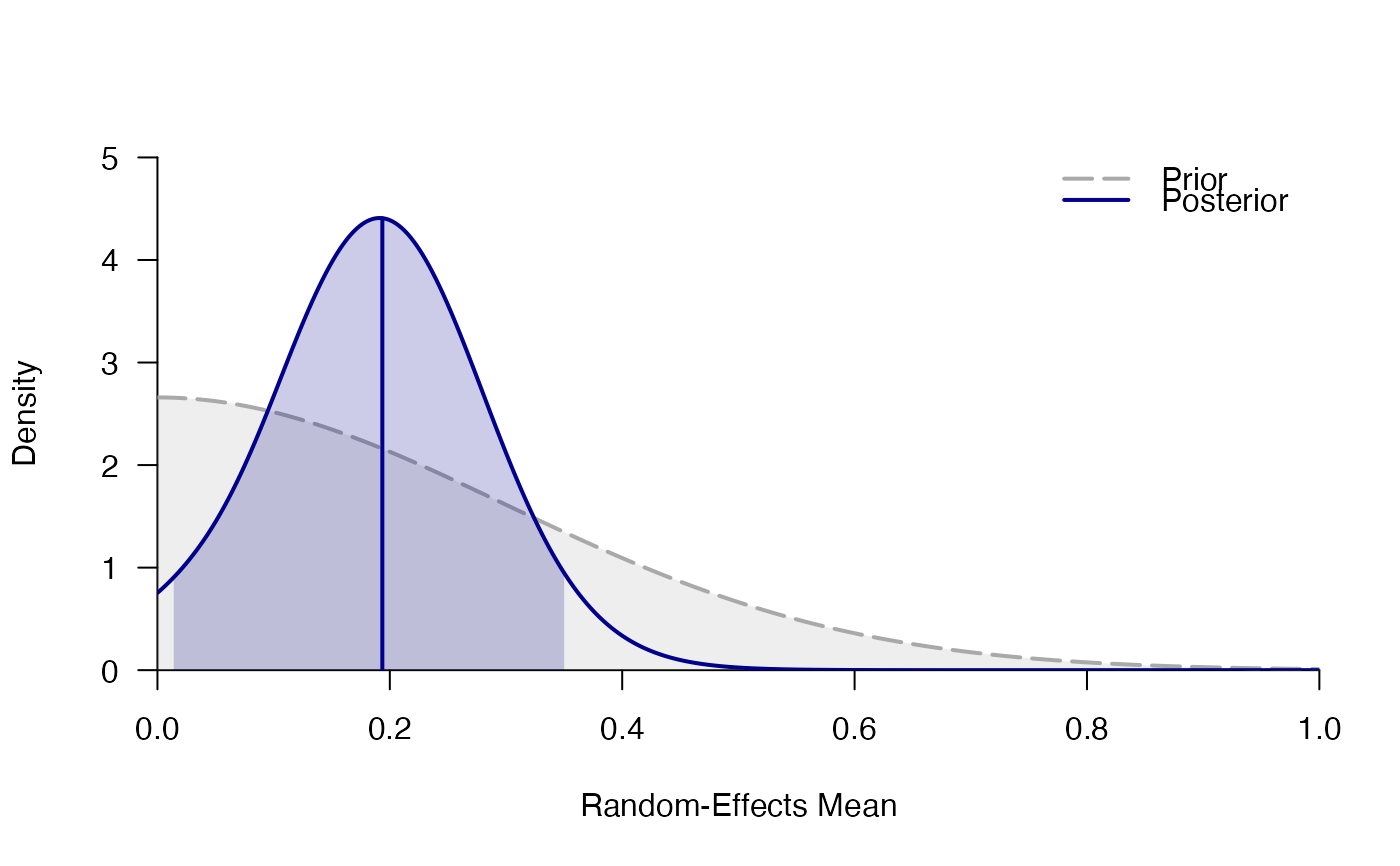 # }
# }